In each simulation a 10% QTL was placed randomly along the genome. (E) The locus-specific power to detect a QTL explaining 10% of the phenotypic variance, from 40,000 simulations. The probability for founder s at locus L is represented by a grey bar at coordinate ( L,s), the shade of grey varying from white ( P = 0) to black ( P = 1). The vertical axis represents the 19 possible founders, s, in alphabetical order. (D) The posterior founder probabilities for ML-100. The vertical grey lines indicate probable recombination breakpoints where the identity of the most probable founder changes. (C) The maximum posterior founder probability, m Li for the ML i = “ML-100”. (B) The maximum posterior founder probability, m Li at locus L, averaged across all MLs i. (A) Number of 10-SNP haplotypes observed among founders (black) and MLs (red) across the genome. thaliana, with vertical red lines marking the chromosome boundaries and the pale blue vertical bars indicating the centromeres. In each panel the x-axis represents the complete 120 Mb genome of A. Our results provide strong support for similar ongoing efforts to produce MAGIC lines in other organisms. We demonstrate the utility of this new mapping population by mapping several known QTL with high precision and by finding novel QTL for germination data and bolting time. We also show how the power to detect a QTL and the mapping accuracy vary, depending on QTL location. We show by simulation that QTL explaining 10% of the phenotypic variance will be detected in most situations with an average mapping error of about 300 kb, and that if the number of lines were doubled the mapping error would be under 200 kb. Analytical methods were developed to fine-map quantitative trait loci (QTL) in the MAGIC lines by reconstructing the genome of each line as a mosaic of the founders. These lines and the 19 founders were genotyped with 1,260 single nucleotide polymorphisms and phenotyped for development-related traits. Here, we present the first panel of MAGIC lines developed: a set of 527 recombinant inbred lines (RILs) descended from a heterogeneous stock of 19 intermated accessions of the plant Arabidopsis thaliana. This approach is expected to improve the precision with which QTL can be mapped, improving the outlook for QTL cloning. Here we describe one such alternative, the Multiparent Advanced Generation Inter-Cross (MAGIC). 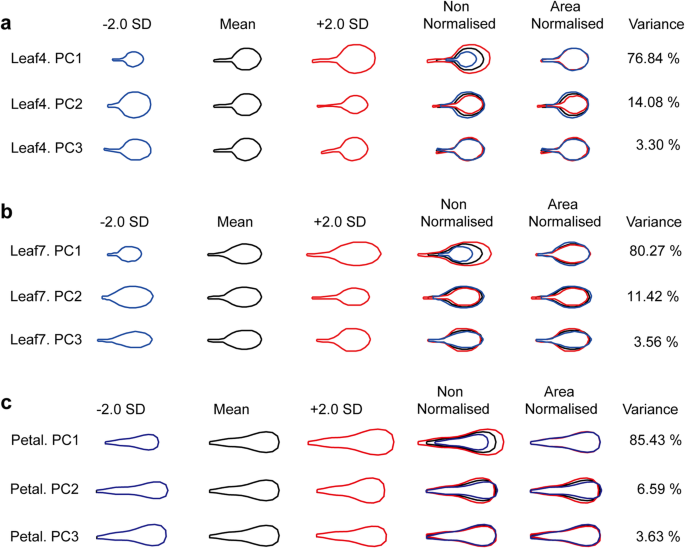
Both of these approaches have some limitations, therefore alternative resources for the genetic dissection of complex traits continue to be sought. Most studies have employed either simple synthetic populations with restricted allelic variation or performed association mapping on a sample of naturally occurring haplotypes. Identifying natural allelic variation that underlies quantitative trait variation remains a fundamental problem in genetics. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
